What is environmental DNA?
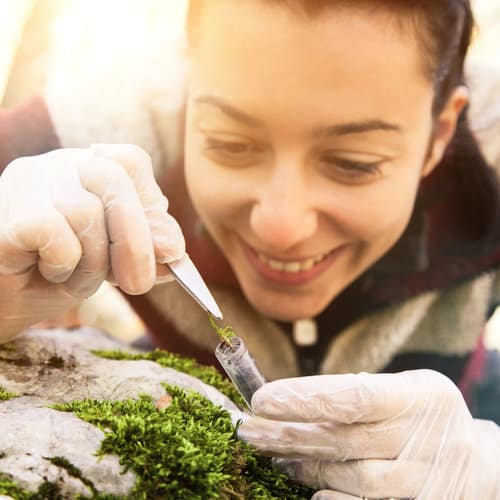
We know that DNA is found in every cell of every living thing, so let’s imagine that a living thing loses a cell. What happens? A plant loses a leaf, a human being scratches himself and loses the superficial cells of his skin, an animal loses its hair, and so on. Living organisms constantly deposit genetic material in their environment.
DNA can be released into the environment via faeces, urine, gametes, mucus, saliva and skin, and it can also come from the decomposition of dead organisms. In aquatic environments, dead, decomposing organisms release a phenomenal amount of genetic material.
So, environmental DNA (eDNA) is the lost cells (intact or not) that we recover from the environment. Scientifically speaking, eDNA is genetic material derived from an environmental sample (freshwater, saltwater, sediment, humus, faeces, etc.).
Why is it useful?
In a context of climate change, it is more than imperative to have a good understanding of the ecosystems that surround us, so that we can better protect them. To implement appropriate strategies and actions to protect our ecosystems, we need to know which species inhabit them. Using sequencing, eDNA identification enables researchers to gain an overview of the community of living beings that inhabit a particular ecosystem, and to deduce trophic links and key species.
Thanks to the eDNA we can extract from these water samples, we’ll have a better idea of the current state of biodiversity in the waterways around us, so we can better understand and protect them.
Mission eDNA : the project
Québec students, a major study of Québec’s waterways is underway, and genomics researchers need your help! Your job is to bring back water samples containing the DNA of various species that live in these ecosystems!
You’ll be working with a professional scientific team to use a powerful and very recent research tool: environmental DNA! It will enable you to recover the traces of DNA that living creatures (fish, invertebrates and microorganisms) leave behind in the water.
With the help of the environmental DNA you’ve collected, we’ll be able to better understand the state of biodiversity in the rivers selected for the project.
This project aims to:
- Contribute to a better understanding of the state of biodiversity in Quebec’s waterways.
- Gather valuable scientific data for the ministère de l’Environnement, de la Lutte contre les changements climatiques, de la Faune et des Parcs (gouv.qc.ca). This data can be used to implement the actions needed to conserve the biodiversity of the targeted watercourses.
- Open a dialogue between researchers from the various ministries involved and students to better understand the influence of human activity on the state of this biodiversity.
The half-day activity will take place on the banks of a river. The results of the analyses will be revealed in the spring, during a live conversation between the students and the scientific team overseeing the project.
This initiative was developed in collaboration with the ministère de l’Environnement, de la Lutte contre les changements climatiques, de la Faune et des Parcs and Université Laval’s Institute of integrative biology and systems. With financial support from the ministère de l’Économie et de l’Innovation.

Related documents
Here are all the documents you need to prepare and run the activity (French only):
- Guide pour le personnel enseignant (Teacher’s guide)
- Cahier de l’élève (Student workbook)
- Grille d’observation (Observation grid)
- Fiche hypothèse (Hypothesis sheet)
- Fiche explicative de la filtration (Filtration explanation sheet)
How-to videos
Here are the three stages of the project (French only):

PHASE 1: Collecting data – Students
This project couldn’t happen without your help! Your students will be the budding researchers who get this project off the ground. You’ll follow a precise research protocol established by a professional scientific team. This will enable you to collect and filter high-quality water samples, from which the eDNA can then be extracted.
You must ensure that your students follow the scientific protocol.
PHASE 2: Analysis – The scientific team
Certifying the health of watercourses
Once the species have been identified, researchers and students will get together to talk about the health of Quebec’s waterways.
Experience overview
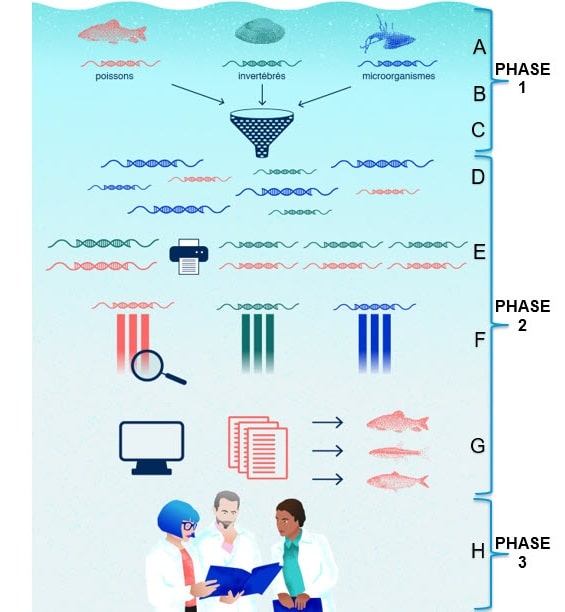
Methodology
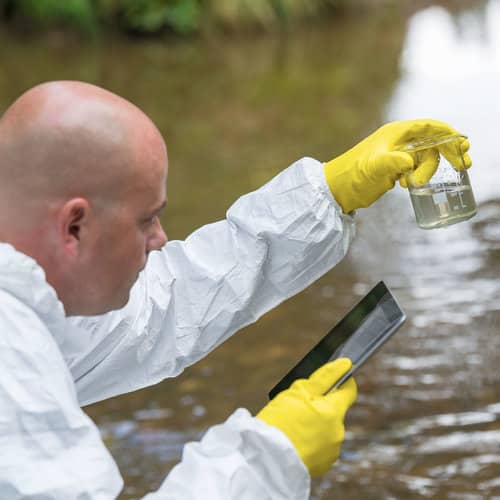
A. Sampling: A water sample is taken from a stream. It contains traces of DNA from fish, invertebrates and microorganisms. This is eDNA.
B. Filtration: The water sample is passed through a fine filter by the students to recover the DNA fragments.
C. Hypothesis: Students are asked to formulate a hypothesis.
Phase 2 : Scientific team
D. Extraction: The DNA is cleaned and preserved, while everything else is discarded (debris, dust, sediment, etc.).
E. Amplification: Our sample contains much more microbial DNA than fish and invertebrate DNA. As in the vast majority of ecosystems, the community of microorganisms in our river, though microscopic, is much larger than that of macroscopic species. The quantities of DNA collected will reflect this difference, and we can therefore expect our sample to contain 90% microbial DNA, compared with 10% DNA from other sources (fish, invertebrates, amphibians, humans, plants, etc.). In order to compensate for the possible biases this may entail during analysis, we amplify (copy in multiple copies) the DNA from fish and invertebrates, since these are the species we also want to study. Amplification takes place using a polymerase chain reaction (PCR). This technique, which takes place over several three-step cycles at different temperatures, produces millions of copies of a DNA molecule using the Taq DNA polymerase enzyme.
F. Sequencing: DNA fragments are read by a sequencer. The affinity of the base pairs is used to decode the order in which they are arranged. To do this, a single-stranded DNA fragment is deposited in a solution containing modified free nitrogenous bases. Luminescent molecules have previously been linked to the various nitrogenous bases. (e.g. Adenine – red, Cytosine – yellow, Thymine – green and Guanine – blue). The modified nitrogenous bases are left to form the DNA strand complementary to our fragment. The nitrogenous bases always associate with the same specificity: adenine with thymine (A-T or T-A) and cytosine with guanine (C-G or G-C). With the sequencer, we can read the “colored” sequence attached to our initial DNA fragment. For a given sequence: red-blue-green-yellow-green-blue-red-red, we can deduce that the sequence of luminescent nitrogenous bases is: A-G-T-C-T-G-A-A and that the initial DNA sequence was: T-C-A-G-A-C-T-T. Record these readings in a computer file.
G. Bioinformatics analysis: Public databases contain the complete genome sequence of the different species studied, so we can compare the sequence we have read with those recorded. So we search the databases of fish, invertebrates and microorganisms for the species to which the DNA we’ve collected and sequenced belongs.
Phase 3: Together
H. Results and interpretation: The researchers analyze the results and try to understand what they mean.
- Why do we find more microorganisms in one river than another?
- Why isn’t such and such a species of fish found in this stream?
- What do these data say about the health of the streams we sampled?